Embedding APIs for RAG: Model Comparison & Implementation Guide (2026)
TL;DR
Embeddings are the backbone of every Retrieval-Augmented Generation system. This guide compares the top embedding models of 2026 — OpenAI text-embedding-3-large, Cohere embed-v4, Voyage 3.5, and BGE-M3 — with benchmarks, pricing, and dimensions. You’ll learn how to generate embeddings, build a complete RAG pipeline in Python, choose the right chunking strategy, pick a vector database, optimize retrieval with hybrid search and reranking, and keep costs under control at scale. Whether you’re building your first RAG prototype or optimizing a production system handling millions of queries, this guide covers what you need.
What Are Embeddings and Why They Matter for RAG
An embedding is a numerical representation of text as a dense vector — a list of floating-point numbers that captures the semantic meaning of a passage. Two pieces of text with similar meanings produce vectors that are close together in the embedding space, even if they use entirely different words.
This property is what makes embeddings essential for RAG. When a user asks a question, you embed the query, search your document embeddings for the most similar vectors, and pass the matching text chunks to a language model as context. The LLM then generates an answer grounded in your actual data rather than relying solely on its training knowledge.
Why RAG Needs Good Embeddings
The quality of your embedding model directly determines the quality of your RAG system. If the embeddings fail to capture the semantic relationship between a user’s query and the relevant document passage, no amount of prompt engineering or LLM quality will save you. Garbage retrieval in, garbage answers out.
Here’s what good embeddings give you:
- Semantic matching: A query like “how to fix memory leaks in Python” should match documents about “debugging Python memory issues” and “garbage collection problems,” not just documents containing the exact phrase “memory leaks.”
- Cross-lingual understanding: Multilingual models can match a question in English with an answer in Japanese, enabling retrieval across language barriers.
- Efficient search: Dense vectors enable approximate nearest neighbor (ANN) search algorithms that can query billions of documents in milliseconds.
- Compact representation: A 500-word paragraph gets compressed into a single vector of 1024-3072 numbers, making storage and comparison computationally tractable.
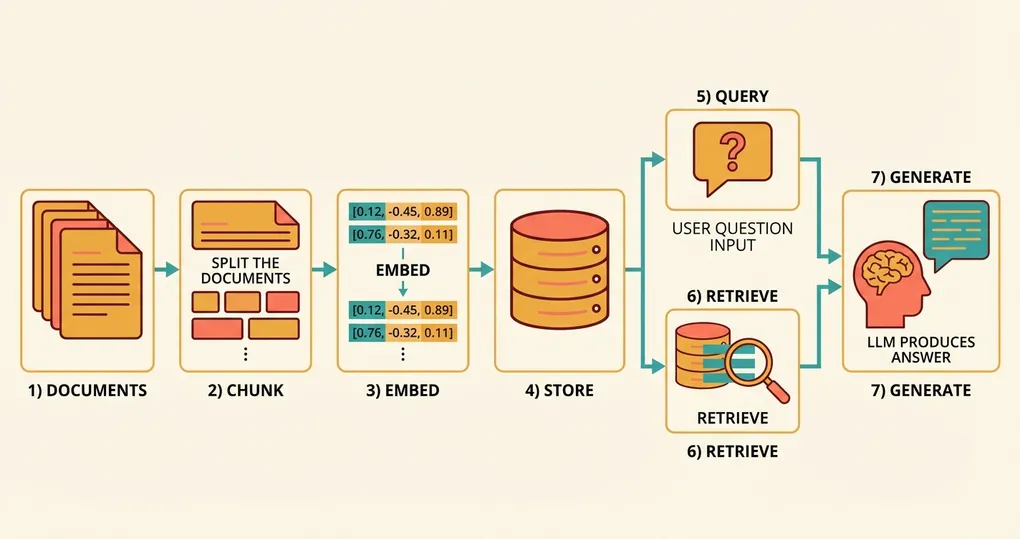
The Embedding-to-Answer Pipeline
At a high level, a RAG system follows this flow:
- Indexing phase (offline): Split your documents into chunks, embed each chunk, store the vectors in a database.
- Query phase (online): Embed the user’s question, search for the nearest vectors, retrieve the original text, pass it to an LLM to generate an answer.
Each step has multiple design decisions that affect accuracy, latency, and cost. We’ll cover all of them.
Top Embedding Models Comparison (2026)
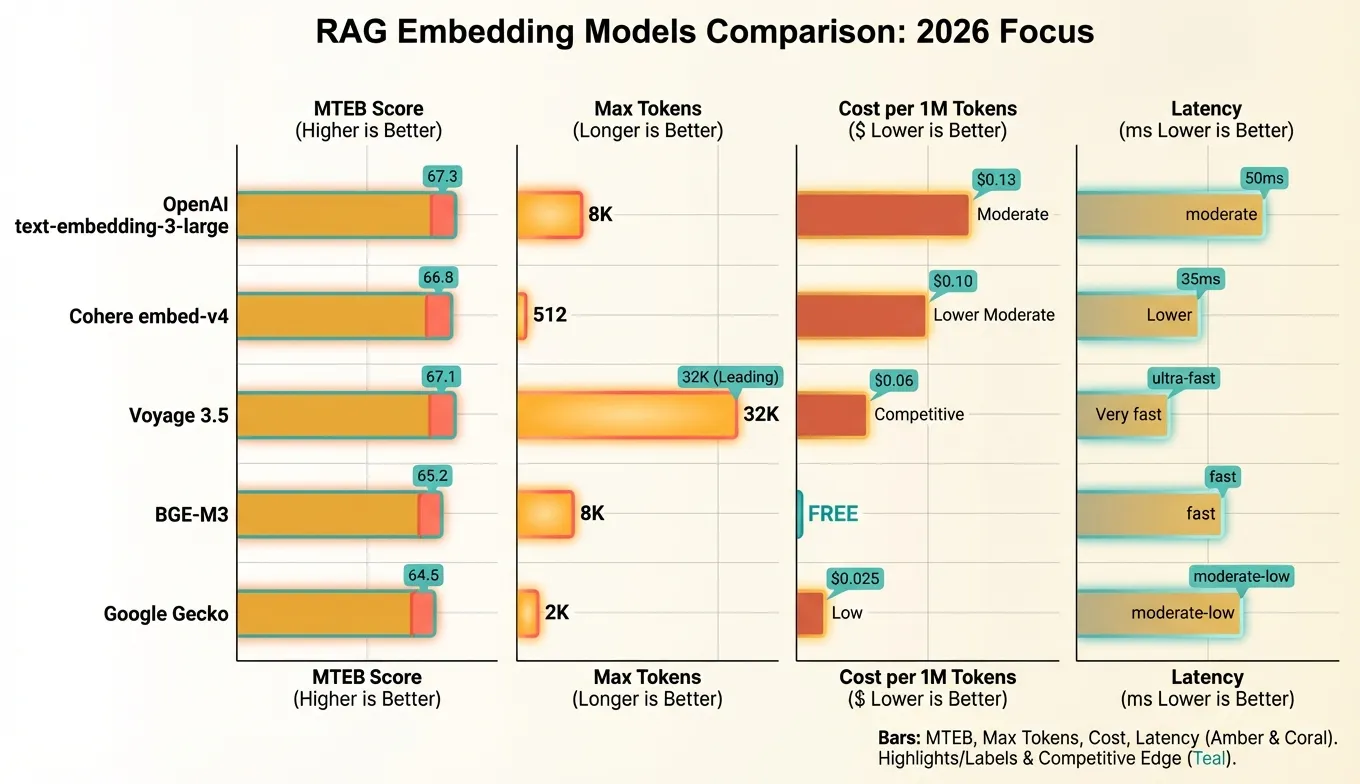
The embedding model landscape has matured considerably. Here’s how the leading options stack up:
| Model | Provider | Dimensions | Max Tokens | MTEB Retrieval Avg | Price (per 1M tokens) | Multilingual | Key Strength |
|---|---|---|---|---|---|---|---|
| text-embedding-3-large | OpenAI | 3072 (configurable) | 8,191 | 67.3 | $0.13 | Limited | Flexible dims, wide adoption |
| embed-v4 | Cohere | 1024 | 512 | 66.8 | $0.10 | 100+ languages | Multilingual, built-in search/doc types |
| voyage-3.5 | Voyage AI | 1024 | 32,000 | 67.1 | $0.06 | Moderate | Long context, code retrieval |
| BGE-M3 | BAAI | 1024 | 8,192 | 65.2 | Free (self-hosted) | 100+ languages | Open source, no API costs |
| Gecko | 768 | 2,048 | 64.5 | $0.025 | Moderate | Low cost, Google ecosystem |
Key Takeaways from the Comparison
OpenAI text-embedding-3-large remains the default choice for many teams. Its configurable dimensionality is unique — you can request 256-dimension vectors for low-cost applications or 3072 for maximum accuracy, all from the same model. The OpenAI SDK is ubiquitous and well-documented.
Cohere embed-v4 is the multilingual champion. If your documents span multiple languages, Cohere’s model handles cross-lingual retrieval natively. It also supports explicit input types (search_document vs search_query), which improves retrieval quality by optimizing the embedding based on its intended use.
Voyage 3.5 stands out for two reasons: an extremely long 32,000-token context window (useful for embedding entire documents without chunking) and excellent code retrieval performance. If you’re building a code search system, Voyage is worth serious consideration. The pricing is also very competitive.
BGE-M3 (from the Beijing Academy of AI) is the leading open-source model. It supports dense, sparse, and multi-vector retrieval in a single model, making it uniquely versatile. Running it yourself requires GPU infrastructure but eliminates per-token costs entirely — a major advantage at scale.
Generating Embeddings with Python
Let’s get practical. Here’s how to generate embeddings using the OpenAI SDK, which is the most common approach and is also compatible with aggregation platforms that expose an OpenAI-compatible endpoint.
Basic Embedding Generation
from openai import OpenAI
client = OpenAI(api_key="your-api-key")
def get_embedding(text: str, model: str = "text-embedding-3-large", dimensions: int = 1024) -> list[float]:
"""Generate an embedding for a single text string."""
response = client.embeddings.create(
input=text,
model=model,
dimensions=dimensions # Only supported by OpenAI's -3 models
)
return response.data[0].embedding
# Example usage
query_embedding = get_embedding("How does photosynthesis work?")
print(f"Embedding dimension: {len(query_embedding)}") # 1024
print(f"First 5 values: {query_embedding[:5]}")Batch Embedding for Efficiency
The API accepts up to 2,048 texts in a single request. Always batch your embeddings to minimize HTTP overhead:
from openai import OpenAI
client = OpenAI(api_key="your-api-key")
def get_embeddings_batch(
texts: list[str],
model: str = "text-embedding-3-large",
dimensions: int = 1024,
batch_size: int = 2048
) -> list[list[float]]:
"""Generate embeddings for a list of texts in batches."""
all_embeddings = []
for i in range(0, len(texts), batch_size):
batch = texts[i : i + batch_size]
response = client.embeddings.create(
input=batch,
model=model,
dimensions=dimensions
)
# Sort by index to maintain original order
batch_embeddings = [item.embedding for item in sorted(response.data, key=lambda x: x.index)]
all_embeddings.extend(batch_embeddings)
return all_embeddings
# Embed 5000 document chunks
chunks = ["Document chunk text here..."] * 5000
embeddings = get_embeddings_batch(chunks)
print(f"Generated {len(embeddings)} embeddings")Using Cohere’s Input Types
Cohere’s API distinguishes between documents and queries, which improves retrieval quality:
import cohere
co = cohere.ClientV2(api_key="your-cohere-key")
# Embed documents (during indexing)
doc_response = co.embed(
texts=["Photosynthesis converts sunlight into chemical energy..."],
model="embed-v4",
input_type="search_document",
embedding_types=["float"]
)
doc_embedding = doc_response.embeddings.float_[0]
# Embed queries (during search)
query_response = co.embed(
texts=["How do plants make food from sunlight?"],
model="embed-v4",
input_type="search_query",
embedding_types=["float"]
)
query_embedding = query_response.embeddings.float_[0]Using an Aggregation Endpoint
If you use an API aggregation platform like Ofox.ai, you can access multiple embedding models through a single OpenAI-compatible endpoint. This simplifies switching between models:
from openai import OpenAI
# Point to the aggregation endpoint
client = OpenAI(
api_key="your-aggregation-key",
base_url="https://api.ofox.ai/v1"
)
# Switch models by changing a single parameter
openai_embedding = client.embeddings.create(
input="test text",
model="text-embedding-3-large",
dimensions=1024
)
# Same client, different model
cohere_embedding = client.embeddings.create(
input="test text",
model="embed-v4"
)Building a RAG Pipeline Step by Step
Let’s build a complete RAG pipeline from scratch. We’ll use OpenAI embeddings, Qdrant as the vector database, and Claude for answer generation.
Step 1: Document Loading
from pathlib import Path
def load_documents(directory: str) -> list[dict]:
"""Load text documents from a directory."""
docs = []
for filepath in Path(directory).glob("**/*.txt"):
text = filepath.read_text(encoding="utf-8")
docs.append({
"id": str(filepath),
"text": text,
"metadata": {
"filename": filepath.name,
"directory": str(filepath.parent)
}
})
return docs
documents = load_documents("./knowledge_base")
print(f"Loaded {len(documents)} documents")Step 2: Chunking
def recursive_chunk(text: str, max_tokens: int = 512, overlap: int = 50) -> list[str]:
"""
Split text into chunks using recursive character splitting.
Uses approximate token count (1 token ≈ 4 characters).
"""
max_chars = max_tokens * 4
overlap_chars = overlap * 4
separators = ["\n\n", "\n", ". ", " "]
def split_text(text: str, separators: list[str]) -> list[str]:
if len(text) <= max_chars:
return [text.strip()] if text.strip() else []
sep = separators[0] if separators else " "
remaining_seps = separators[1:] if len(separators) > 1 else separators
parts = text.split(sep)
chunks = []
current_chunk = ""
for part in parts:
candidate = f"{current_chunk}{sep}{part}" if current_chunk else part
if len(candidate) <= max_chars:
current_chunk = candidate
else:
if current_chunk:
chunks.append(current_chunk.strip())
# If a single part is too long, split it further
if len(part) > max_chars and remaining_seps:
chunks.extend(split_text(part, remaining_seps))
else:
current_chunk = part
if current_chunk:
chunks.append(current_chunk.strip())
return chunks
raw_chunks = split_text(text, separators)
# Add overlap between consecutive chunks
if overlap_chars > 0 and len(raw_chunks) > 1:
overlapped = [raw_chunks[0]]
for i in range(1, len(raw_chunks)):
prev_tail = raw_chunks[i - 1][-overlap_chars:]
overlapped.append(prev_tail + " " + raw_chunks[i])
return overlapped
return raw_chunksStep 3: Embed and Store
from openai import OpenAI
from qdrant_client import QdrantClient
from qdrant_client.models import Distance, VectorParams, PointStruct
import uuid
# Initialize clients
openai_client = OpenAI(api_key="your-api-key")
qdrant = QdrantClient(url="http://localhost:6333")
COLLECTION_NAME = "knowledge_base"
EMBEDDING_DIM = 1024
# Create collection
qdrant.create_collection(
collection_name=COLLECTION_NAME,
vectors_config=VectorParams(
size=EMBEDDING_DIM,
distance=Distance.COSINE
)
)
def index_documents(documents: list[dict]) -> int:
"""Chunk, embed, and store documents in Qdrant."""
points = []
for doc in documents:
chunks = recursive_chunk(doc["text"])
if not chunks:
continue
# Batch embed all chunks for this document
response = openai_client.embeddings.create(
input=chunks,
model="text-embedding-3-large",
dimensions=EMBEDDING_DIM
)
for j, (chunk, item) in enumerate(zip(chunks, sorted(response.data, key=lambda x: x.index))):
points.append(PointStruct(
id=str(uuid.uuid4()),
vector=item.embedding,
payload={
"text": chunk,
"doc_id": doc["id"],
"chunk_index": j,
**doc["metadata"]
}
))
# Upsert in batches of 100
for i in range(0, len(points), 100):
qdrant.upsert(
collection_name=COLLECTION_NAME,
points=points[i : i + 100]
)
return len(points)
total_chunks = index_documents(documents)
print(f"Indexed {total_chunks} chunks")Step 4: Retrieval and Answer Generation
from openai import OpenAI
import anthropic
openai_client = OpenAI(api_key="your-openai-key")
claude_client = anthropic.Anthropic(api_key="your-anthropic-key")
def retrieve(query: str, top_k: int = 5) -> list[dict]:
"""Retrieve the most relevant chunks for a query."""
query_response = openai_client.embeddings.create(
input=query,
model="text-embedding-3-large",
dimensions=EMBEDDING_DIM
)
query_vector = query_response.data[0].embedding
results = qdrant.query_points(
collection_name=COLLECTION_NAME,
query=query_vector,
limit=top_k,
with_payload=True
)
return [
{"text": point.payload["text"], "score": point.score, "doc_id": point.payload["doc_id"]}
for point in results.points
]
def rag_answer(question: str) -> str:
"""Full RAG pipeline: retrieve context, then generate an answer."""
# Retrieve relevant chunks
chunks = retrieve(question, top_k=5)
# Format context
context = "\n\n---\n\n".join(
f"[Source: {c['doc_id']} | Relevance: {c['score']:.3f}]\n{c['text']}"
for c in chunks
)
# Generate answer with Claude
response = claude_client.messages.create(
model="claude-sonnet-4-6-20260301",
max_tokens=2000,
system="You are a helpful assistant. Answer questions based on the provided context. If the context doesn't contain enough information, say so clearly. Cite sources when possible.",
messages=[{
"role": "user",
"content": f"Context:\n{context}\n\nQuestion: {question}\n\nAnswer based on the context above:"
}]
)
return response.content[0].text
# Use it
answer = rag_answer("What are the main causes of climate change?")
print(answer)Chunking Strategies: Choosing the Right Approach
The way you split documents into chunks has a surprisingly large impact on RAG quality. Poor chunking leads to either incomplete context (chunks too small) or diluted relevance (chunks too large). Here are the three main approaches.
Fixed-Size Chunking
The simplest strategy: split text into chunks of N tokens with M tokens of overlap.
Pros: Fast, deterministic, easy to implement. Cons: Splits mid-sentence and mid-paragraph, breaking semantic coherence. Best for: Homogeneous content like log files, CSV data, or simple text where structure doesn’t matter.
import tiktoken
def fixed_size_chunk(text: str, chunk_size: int = 512, overlap: int = 50) -> list[str]:
"""Split text into fixed-size token chunks with overlap."""
enc = tiktoken.encoding_for_model("gpt-4")
tokens = enc.encode(text)
chunks = []
start = 0
while start < len(tokens):
end = start + chunk_size
chunk_tokens = tokens[start:end]
chunks.append(enc.decode(chunk_tokens))
start = end - overlap
return chunksRecursive Character Splitting
The standard approach used by LangChain and most RAG frameworks. It tries to split on paragraph boundaries first, then sentences, then words — preserving as much semantic structure as possible.
Pros: Respects document structure, produces coherent chunks. Cons: Chunk sizes vary, may produce very small chunks from short sections. Best for: General-purpose text, articles, documentation.
This is the strategy we implemented in Step 2 above. It’s the recommended default for most use cases.
Semantic Chunking
The most sophisticated approach: use embeddings to detect topic shifts and place chunk boundaries where the topic changes.
Pros: Chunks are topically coherent, leading to better retrieval precision. Cons: Requires an embedding model pass during chunking (added cost and complexity), harder to debug. Best for: Long documents with multiple topics, legal documents, research papers.
import numpy as np
from openai import OpenAI
client = OpenAI(api_key="your-api-key")
def semantic_chunk(text: str, threshold: float = 0.3) -> list[str]:
"""Split text into chunks based on semantic similarity between sentences."""
# Split into sentences
sentences = [s.strip() for s in text.replace("\n", " ").split(". ") if s.strip()]
if len(sentences) <= 1:
return [text]
# Embed all sentences
response = client.embeddings.create(
input=sentences,
model="text-embedding-3-large",
dimensions=256 # Low dims is fine for similarity comparison
)
embeddings = [item.embedding for item in sorted(response.data, key=lambda x: x.index)]
# Find breakpoints where consecutive sentences are dissimilar
breakpoints = [0]
for i in range(1, len(embeddings)):
similarity = np.dot(embeddings[i - 1], embeddings[i]) / (
np.linalg.norm(embeddings[i - 1]) * np.linalg.norm(embeddings[i])
)
if similarity < (1 - threshold): # Low similarity = topic shift
breakpoints.append(i)
breakpoints.append(len(sentences))
# Group sentences into chunks
chunks = []
for i in range(len(breakpoints) - 1):
chunk_sentences = sentences[breakpoints[i]:breakpoints[i + 1]]
chunks.append(". ".join(chunk_sentences) + ".")
return chunksRecommended Chunk Size Guidelines
| Content Type | Chunk Size (tokens) | Overlap (tokens) | Strategy |
|---|---|---|---|
| General articles | 512 | 50 | Recursive |
| Technical documentation | 384 | 64 | Recursive |
| Legal documents | 768 | 100 | Semantic |
| Code files | Function-level | 0 | AST-aware |
| FAQ/Q&A pairs | Per question | 0 | Fixed (by item) |
| Chat transcripts | Per turn or group | 1 turn | Fixed (by turn) |
Vector Database Options
Your vector database stores the embeddings and performs similarity search. Here’s how the major options compare for production RAG systems.
Comparison Table
| Database | Type | Max Vectors | Hybrid Search | Filtering | Pricing Model | Best For |
|---|---|---|---|---|---|---|
| Pinecone | Managed | Billions | Yes (2024+) | Metadata | Per pod/serverless | Teams wanting zero ops |
| Qdrant | Self-hosted / Cloud | Billions | Yes | Payload filters | Free (self-hosted) / Usage | Performance-critical apps |
| Weaviate | Self-hosted / Cloud | Billions | Yes (BM25 + vector) | GraphQL filters | Free (self-hosted) / Usage | Complex filtering needs |
| pgvector | PostgreSQL extension | Millions | With pg_search | Full SQL | Free (self-hosted) | Teams already on PostgreSQL |
| Chroma | Embedded | Millions | No | Metadata | Free (open source) | Prototyping, small datasets |
| Milvus | Self-hosted / Cloud | Billions | Yes | Attribute filtering | Free (self-hosted) / Zilliz Cloud | Large-scale deployment |
pgvector: Minimal Infrastructure
If you already use PostgreSQL, pgvector is the lowest-friction option:
import psycopg2
from pgvector.psycopg2 import register_vector
conn = psycopg2.connect("postgresql://localhost/mydb")
register_vector(conn)
cur = conn.cursor()
# Create table with vector column
cur.execute("""
CREATE TABLE IF NOT EXISTS documents (
id SERIAL PRIMARY KEY,
text TEXT NOT NULL,
metadata JSONB,
embedding vector(1024)
)
""")
# Create HNSW index for fast search
cur.execute("""
CREATE INDEX IF NOT EXISTS docs_embedding_idx
ON documents USING hnsw (embedding vector_cosine_ops)
WITH (m = 16, ef_construction = 200)
""")
conn.commit()
# Insert a document
cur.execute(
"INSERT INTO documents (text, metadata, embedding) VALUES (%s, %s, %s)",
("Example document text", '{"source": "wiki"}', embedding_vector)
)
conn.commit()
# Search
cur.execute(
"SELECT text, metadata, 1 - (embedding <=> %s::vector) AS similarity FROM documents ORDER BY embedding <=> %s::vector LIMIT 5",
(query_vector, query_vector)
)
results = cur.fetchall()Qdrant: High Performance
Qdrant offers excellent single-node performance and a clean Python client:
from qdrant_client import QdrantClient
from qdrant_client.models import Distance, VectorParams, PointStruct, Filter, FieldCondition, MatchValue
client = QdrantClient(url="http://localhost:6333")
# Create collection
client.create_collection(
collection_name="docs",
vectors_config=VectorParams(size=1024, distance=Distance.COSINE)
)
# Search with metadata filtering
results = client.query_points(
collection_name="docs",
query=query_vector,
query_filter=Filter(
must=[FieldCondition(key="category", match=MatchValue(value="engineering"))]
),
limit=10
)Choosing the Right Database
For prototyping: Start with Chroma (embedded, zero setup) or pgvector (if you already have PostgreSQL).
For production under 10M vectors: pgvector with HNSW indexes handles this comfortably. One less service to manage.
For production over 10M vectors: Qdrant or Pinecone. You’ll need dedicated vector search infrastructure at this scale, and these are battle-tested.
For hybrid search as a priority: Weaviate has the most mature hybrid search (BM25 + vector), with fine-grained control over the weighting between keyword and semantic results.
Retrieval Optimization: Hybrid Search and Reranking
Vector similarity search alone has a well-known weakness: it can miss results that are lexically relevant but semantically distant. A user searching for “error code E-4012” might not find the document about that specific error if the embedding model doesn’t capture the exact code. This is where hybrid search and reranking come in.
Hybrid Search
Hybrid search combines two retrieval methods:
- Dense retrieval (vector similarity): Finds semantically similar documents.
- Sparse retrieval (BM25/keyword matching): Finds documents with matching keywords.
The results are merged using Reciprocal Rank Fusion (RRF) or a weighted combination:
def hybrid_search(query: str, alpha: float = 0.7, top_k: int = 10) -> list[dict]:
"""
Combine dense and sparse search results.
alpha: weight for dense search (1.0 = pure vector, 0.0 = pure keyword)
"""
# Dense search
query_embedding = get_embedding(query)
dense_results = vector_db.search(query_embedding, limit=top_k * 2)
# Sparse search (BM25)
sparse_results = bm25_index.search(query, limit=top_k * 2)
# Reciprocal Rank Fusion
scores = {}
k = 60 # RRF constant
for rank, result in enumerate(dense_results):
doc_id = result["id"]
scores[doc_id] = scores.get(doc_id, 0) + alpha * (1 / (k + rank + 1))
for rank, result in enumerate(sparse_results):
doc_id = result["id"]
scores[doc_id] = scores.get(doc_id, 0) + (1 - alpha) * (1 / (k + rank + 1))
# Sort by combined score and return top_k
ranked = sorted(scores.items(), key=lambda x: x[1], reverse=True)[:top_k]
return [{"id": doc_id, "score": score} for doc_id, score in ranked]Reranking
Reranking adds a second stage that uses a cross-encoder model to score each retrieved chunk against the query. Cross-encoders are more accurate than bi-encoders (embedding models) because they process the query and document together, capturing fine-grained interactions.
import cohere
co = cohere.ClientV2(api_key="your-cohere-key")
def rerank_results(query: str, documents: list[str], top_n: int = 5) -> list[dict]:
"""Rerank retrieved documents using Cohere Rerank."""
response = co.rerank(
query=query,
documents=documents,
model="rerank-v3.5",
top_n=top_n
)
return [
{"index": result.index, "text": documents[result.index], "relevance_score": result.relevance_score}
for result in response.results
]
# Usage in the RAG pipeline
initial_chunks = retrieve(query, top_k=25) # Broad initial retrieval
chunk_texts = [c["text"] for c in initial_chunks]
reranked = rerank_results(query, chunk_texts, top_n=5) # Narrow to best 5The Retrieve-then-Rerank Pattern
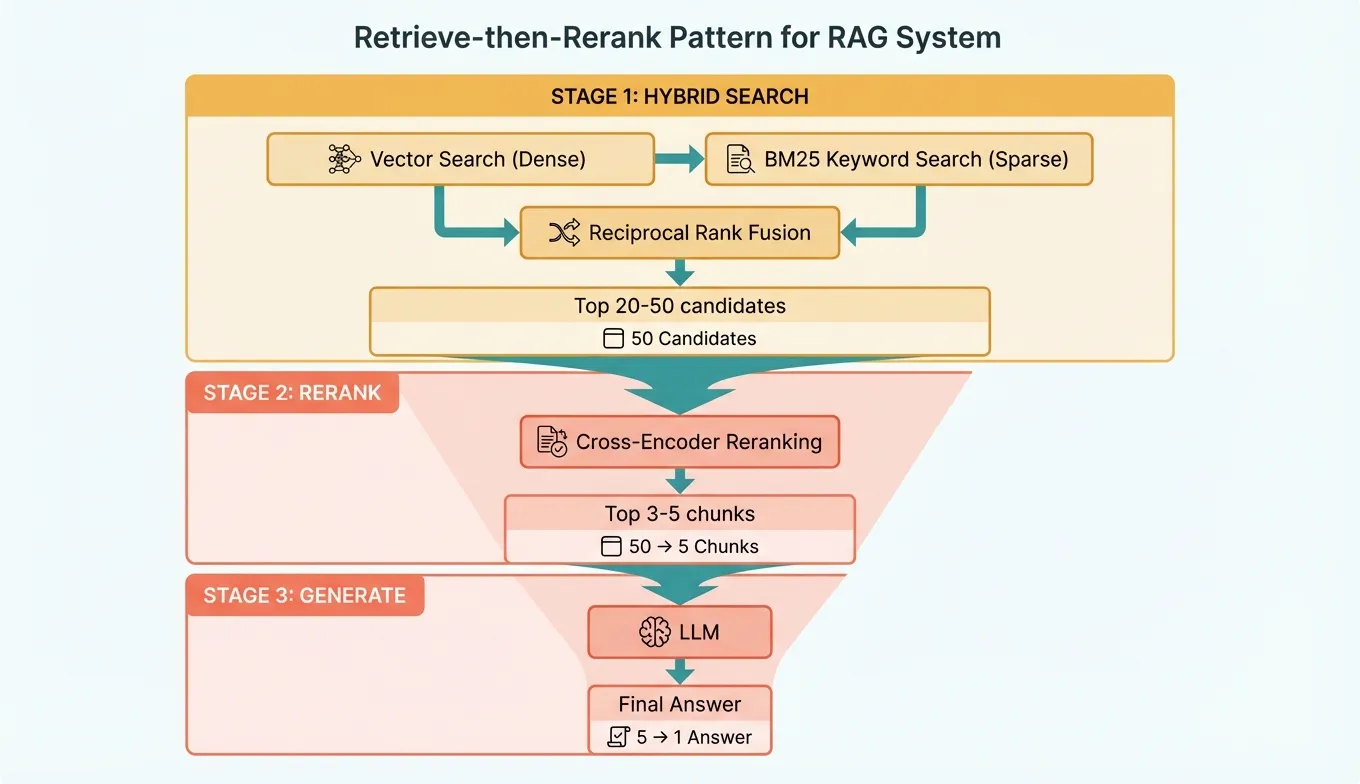
The optimal retrieval pipeline for production RAG follows this pattern:
- Broad retrieval: Fetch top 20-50 candidates using hybrid search (fast, recall-optimized).
- Rerank: Score all candidates with a cross-encoder, select top 3-5 (slow but precise).
- Generate: Pass the top chunks to the LLM for answer generation.
This two-stage approach gives you the best of both worlds: high recall from the broad first stage and high precision from the reranking second stage.
Cost Optimization for Embeddings at Scale
Embedding costs can add up quickly when you’re processing millions of documents. Here are concrete strategies to keep costs manageable.
1. Reduce Dimensions
OpenAI’s text-embedding-3-large supports native dimension reduction. Dropping from 3072 to 1024 dimensions:
- Reduces storage by 66%
- Reduces vector search latency by approximately 50%
- Costs the same per API call (pricing is per token, not per dimension)
- Only loses 1-2% retrieval accuracy on most benchmarks
Always use the dimensions parameter — there’s rarely a reason to store full 3072-dimension vectors.
2. Deduplicate Before Embedding
Don’t embed duplicate or near-duplicate content. Hash your chunks and skip duplicates:
import hashlib
def deduplicate_chunks(chunks: list[str]) -> list[str]:
"""Remove exact duplicate chunks."""
seen = set()
unique = []
for chunk in chunks:
h = hashlib.sha256(chunk.encode()).hexdigest()
if h not in seen:
seen.add(h)
unique.append(chunk)
return uniqueFor large-scale deduplication, use MinHash or SimHash to also catch near-duplicates (chunks that differ by only a few words).
3. Use Open-Source Models at Scale
Beyond approximately 50 million tokens of embedding volume per month, self-hosting BGE-M3 on a GPU becomes cheaper than API calls:
| Monthly Volume | OpenAI text-embedding-3-large | Self-hosted BGE-M3 (A100) |
|---|---|---|
| 10M tokens | $1.30 | ~$15 (over-provisioned) |
| 100M tokens | $13.00 | ~$15 |
| 1B tokens | $130.00 | ~$25 (scales well) |
| 10B tokens | $1,300.00 | ~$100 (multiple GPUs) |
The crossover point is roughly 100-200M tokens per month, depending on your GPU costs.
4. Cache Embeddings Aggressively
Never embed the same text twice. Implement an embedding cache:
import hashlib
import json
from pathlib import Path
class EmbeddingCache:
"""Simple file-based embedding cache."""
def __init__(self, cache_dir: str = ".embedding_cache"):
self.cache_dir = Path(cache_dir)
self.cache_dir.mkdir(exist_ok=True)
def _key(self, text: str, model: str, dims: int) -> str:
raw = f"{model}:{dims}:{text}"
return hashlib.sha256(raw.encode()).hexdigest()
def get(self, text: str, model: str, dims: int) -> list[float] | None:
path = self.cache_dir / f"{self._key(text, model, dims)}.json"
if path.exists():
return json.loads(path.read_text())
return None
def set(self, text: str, model: str, dims: int, embedding: list[float]) -> None:
path = self.cache_dir / f"{self._key(text, model, dims)}.json"
path.write_text(json.dumps(embedding))For production, replace the file-based cache with Redis or another key-value store.
5. Batch and Parallelize
Always embed in batches. The difference between 1,000 individual API calls and a single batch call with 1,000 texts is enormous — not in token cost, but in HTTP overhead, rate limiting, and wall-clock time.
import asyncio
from openai import AsyncOpenAI
async_client = AsyncOpenAI(api_key="your-api-key")
async def embed_all_chunks(chunks: list[str], batch_size: int = 2048) -> list[list[float]]:
"""Embed all chunks with concurrent batch requests."""
batches = [chunks[i:i + batch_size] for i in range(0, len(chunks), batch_size)]
async def embed_batch(batch: list[str]) -> list[list[float]]:
response = await async_client.embeddings.create(
input=batch,
model="text-embedding-3-large",
dimensions=1024
)
return [item.embedding for item in sorted(response.data, key=lambda x: x.index)]
# Run up to 5 batches concurrently
semaphore = asyncio.Semaphore(5)
async def limited_embed(batch: list[str]) -> list[list[float]]:
async with semaphore:
return await embed_batch(batch)
results = await asyncio.gather(*[limited_embed(b) for b in batches])
return [emb for batch_result in results for emb in batch_result]Common Pitfalls and How to Avoid Them
Building RAG systems involves many subtle decisions. Here are the mistakes that trip up most teams.
1. Chunks Too Large or Too Small
Problem: Chunks of 2000+ tokens dilute the relevant information with noise, reducing retrieval precision. Chunks under 100 tokens lack enough context for the LLM to generate good answers.
Fix: Start with 512 tokens and 50 tokens of overlap. Measure retrieval recall on a test set and adjust. If recall is low, try smaller chunks (256-384). If the LLM struggles with incomplete context, try larger chunks (768-1024) or retrieve more chunks.
2. Not Normalizing Text Before Embedding
Problem: Extra whitespace, inconsistent encoding, HTML artifacts, and other noise reduce embedding quality.
Fix: Clean text before embedding:
import re
import unicodedata
def normalize_text(text: str) -> str:
"""Normalize text before embedding."""
text = unicodedata.normalize("NFKC", text)
text = re.sub(r"\s+", " ", text) # Collapse whitespace
text = re.sub(r"<[^>]+>", "", text) # Strip HTML
text = text.strip()
return text3. Using the Wrong Distance Metric
Problem: Using Euclidean distance when the model was trained for cosine similarity (or vice versa) degrades results.
Fix: Use cosine similarity for OpenAI and Cohere models. Check the model documentation for the recommended metric. Most modern embedding models are trained with cosine similarity in mind.
4. Ignoring the Query-Document Asymmetry
Problem: User queries are short (“how to fix memory leak”), while documents are long paragraphs. Embedding both the same way loses information.
Fix: Use models that support input types (Cohere’s search_query vs search_document). For models without explicit types, consider prepending “search query: ” to queries and “search document: ” to documents — this is a technique from the E5 model family that often helps even with other models.
5. Not Evaluating Retrieval Quality Separately from Generation Quality
Problem: When the RAG system gives bad answers, teams blame the LLM. Often, the retrieval step is the real problem — the right chunks were never found.
Fix: Evaluate retrieval and generation independently. Build a test set of queries with known relevant chunks. Measure retrieval recall@k (what percentage of relevant chunks appear in the top-k results). Only after retrieval quality is solid should you optimize the generation step.
6. Embedding Stale or Redundant Content
Problem: Re-embedding your entire corpus every time a few documents change wastes money and time.
Fix: Track document hashes. Only re-embed documents whose content has changed. Use your vector database’s upsert functionality to update individual vectors without rebuilding the entire index.
7. Skipping Metadata Filtering
Problem: Searching your entire vector space when the query clearly relates to a specific category, date range, or document type wastes computation and returns irrelevant results.
Fix: Attach metadata (category, date, source, language) to every chunk during indexing. Apply metadata filters before vector search to narrow the search space. This dramatically improves both speed and relevance.
8. Not Planning for Embedding Model Upgrades
Problem: When a better embedding model comes out, you have to re-embed everything — which is expensive and time-consuming.
Fix: Design your system to handle re-embedding. Keep the original text alongside the vectors. Use collection versioning (e.g., docs_v1, docs_v2) so you can run old and new embeddings in parallel during migration. API aggregation platforms like Ofox.ai make it easier to switch between embedding providers since you only need to change the model parameter, not the entire integration.
9. Overcomplicating the Pipeline
Problem: Teams add every optimization at once — hybrid search, reranking, query expansion, hypothetical document embeddings (HyDE) — creating a system that’s complex, slow, and hard to debug.
Fix: Start simple. Basic chunking + vector search + a good LLM will get you surprisingly far. Add complexity only when you’ve measured a specific problem. Profile your pipeline to identify the actual bottleneck before adding more stages.
10. Ignoring Latency Budgets
Problem: A RAG pipeline with embedding generation (100ms) + vector search (50ms) + reranking (200ms) + LLM generation (2000ms) adds up to 2.3+ seconds. For real-time applications, this may be too slow.
Fix: Set a latency budget upfront. If you need sub-second responses, skip reranking and use a faster LLM. Pre-compute query embeddings where possible. Use approximate nearest neighbor search with higher speed settings (lower accuracy is usually acceptable). Consider caching frequent queries and their answers.
Conclusion
Building an effective RAG system is about making smart trade-offs at every layer — from the embedding model you choose, to how you chunk documents, to whether you invest in hybrid search and reranking. The good news is that the tooling has matured enormously. In 2026, you can go from zero to a production-quality RAG pipeline in a weekend, thanks to excellent embedding APIs, robust vector databases, and clear best practices.
Start with the defaults: OpenAI text-embedding-3-large at 1024 dimensions, recursive chunking at 512 tokens, and your vector database of choice. Measure retrieval quality on a representative test set. Then add complexity — hybrid search, reranking, semantic chunking — only where measurements show it’s needed. The simplest pipeline that meets your accuracy and latency requirements is the best pipeline.